Marginal Revolution linked a post at Genomes Unzipped, "Size matters, and other lessons from medical genetics", with the interesting centerpiece graph:

This is from pg 3 of an Ioannidis 2001 et al article (who else?) on what is called a funnel plot: each line represents a series of studies about some particularly hot gene-disease correlations, plotted where Y = the odds ratio (measure of effect size; all results are 'statistically significant', of course) and X = the sample size. The 1 line is the null hypothesis, here. You will notice something dramatic: as we move along the X-axis and sample sizes increase, everything begins to converge on 1:
Readers familiar with the history of medical association studies will be unsurprised by what happened over the next few years: initial excitement (this same polymorphism was associated with diabetes! And longevity!) was followed by inconclusive replication studies and, ultimately, disappointment. In 2000, 8 years after the initial report, a large study involving over 5,000 cases and controls found absolutely no detectable effect of the ACE polymorphism on heart attack risk. In the meantime, the same polymorphism had turned up in dozens of other association studies for a wide range of traits ranging from obstetric cholestasis to meningococcal disease in children, virtually none of which have ever been convincingly replicated.
(See also "Why epidemiology will not correct itself" or the DNB FAQ.)
This graph is interesting as it shows 8 different regressions to the mean. What is also interesting is what a funnel plot is usually used for, why I ran into it in the first place reading Cochrane Group materials - it's used to show publication bias.
That is, suppose you were looking at a gene you know for certain not to be correlated (you knew the null result to be true), and you ran many trials, each with a different number of samples; you would expect that the trials with small samples would have a wide scattering of results (sometimes the effect size would look wildly large and sometimes they would look wildly small or negative), and that this scattering would be equally for and against any connection (on either side of the 1 line). By the same reasoning you would expect that your largest samples would only be scattered a little bit on either side of the 1 line, and the larger the sample the closer they will be to the 1/null line.
If you plotted your hypothetical trials on the above graph, you'd see what looks pretty much like the above graph - a kind of triangular cloud, wide on the left and ever narrowing towards the right as sample sizes increase and variance decreases.
Now here's the question: given that all 8 correlations trend steadily towards the null hypothesis, one would seem to expect them to actually be the null result. But if that is so, where are the random trials scattered on the other side of the 1 line? Not one sequence of studies ever crosses the 1 line!
The ACE story is not unique; time and time again, initial reports of associations between candidate genes and complex diseases failed to replicate in subsequent studies. With the benefit of hindsight, the problem is clear: in general, common genetic polymorphisms have very small effects on disease risk. Detecting these subtle effects requires studying not dozens or hundreds, but thousands or tens-of-thousands of individuals. Smaller studies, which had no power to detect these small effects, were essentially random p-value generators. Sometimes the p-values were “significant” and sometimes not, without any correlation to whether a variant was truly associated. Additionally, since investigators were often looking at only a few variants (often just one!) in a single gene that they strongly believed to be involved in the disease, they were often able to subset the data (splitting males and females, for example) to find “significant” results in some subgroup. This, combined with a tendency to publish positive results and leave negative results in a desk drawer, resulted in a conflicted and confusing body of literature which actively retarded medical genetics progress.
Wikipedia's funnel chart graph shows us how a plot should look (with this time sample size being the Y axis and odds being the X axis, so the triangle is rotated):
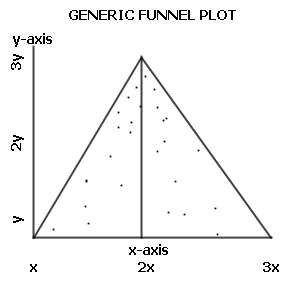
Does that describe any of the sequences graphed above?
I don't think your point applies to this specific graph, since it is a cumulative odds ratio graph selected for initially high error in one direction.
Associations from Ioannides' analysis were selected for inclusion on this graph because there were dramatic changes from the first study to the next studies - in this case, because the first study had a high effect size and the others showed lower effect sizes. Studies were selected for Ioannides' analysis in the first place because there were meta-analyses around them - which means lots of people tried to replicate them - which means the initial result must have been surprising and interesting. So these have been double-selected already for original studies likely to have high errors.
In a cumulative odds ratio graph (unlike the individual odds ratio graphs of the classic funnel plot), most of the work of subsequent studies will go to bringing the trend line closer to the mean. Even a study that shows an effect in the opposite direction as the original won't move the trend line to the other side of the identity line if the effect is smaller. So if the graph is arranged so that the most surprising and deviant result is the first, then it could very well look like this one even if there were no publication bias.
This is doubly true when the graphs are not actually converging to one - Ioannides' paper admits that three of these are probably real associations, and this is most obvious here on the DRD2 line, which converges around .5.
The original article should be edited to include note of this. As it stands, it is misleading. The original plot may or may not show problems, but expecting it to look like the "generic funnel plot" image is wrong.